

Design BAC modification strategy
Once you tell us the changes you wish to make to your BAC, our skilled scientists will design a detailed strategy to achieve the modification. This can be as simple as telling us "I want to put a lacZ reporter after the promoter of this gene."
Leave BAC ordering to us
Too much hassle ordering BACs? Just give us the ID number of the BAC you want, and we can source it for you. The great majority of BACs are available from the BACPAC Resource Center at CHORI, but some BACs have to be obtained from other suppliers. This step is not needed if you send us your own BAC in E. coli.
One-step BAC modification
This modification can be used to introduce either major sequence changes such as placing a reporter behind a gene’s promoter to track gene expression (Figure 1), or minor changes such as point mutations (Figure 2).
In the case of a major change (Figure 1), a DNA fragment containing your gene of interest (GOI) such as lacZ or GFP reporter, plus a drug-selection cassette flanked by FRT sites, along with flanking homology regions, is used to replace an endogenous region on the BAC via homologous recombination. The drug-selection cassette facilitates the identification of successfully modified BAC clones. This cassette can then be removed by Flp-mediate recombination, leaving behind one FRT site.
In the case of a point mutation (Figure 2), a DNA fragment containing the mutated region, plus a drug-selection cassette flanked by FRT sites, along with flanking homology regions, is used to replace the corresponding endogenous region via homologous recombination. This cassette can then be removed by Flp-mediate recombination, leaving behind one FRT site. This leftover FRT is considered a “scar” sequence because the BAC now carries, in addition to the desired point mutation, also this leftover FRT. In order not to affect gene function, this scar sequence should be placed in a nonfunctional region of the gene, such as intron or UTR.
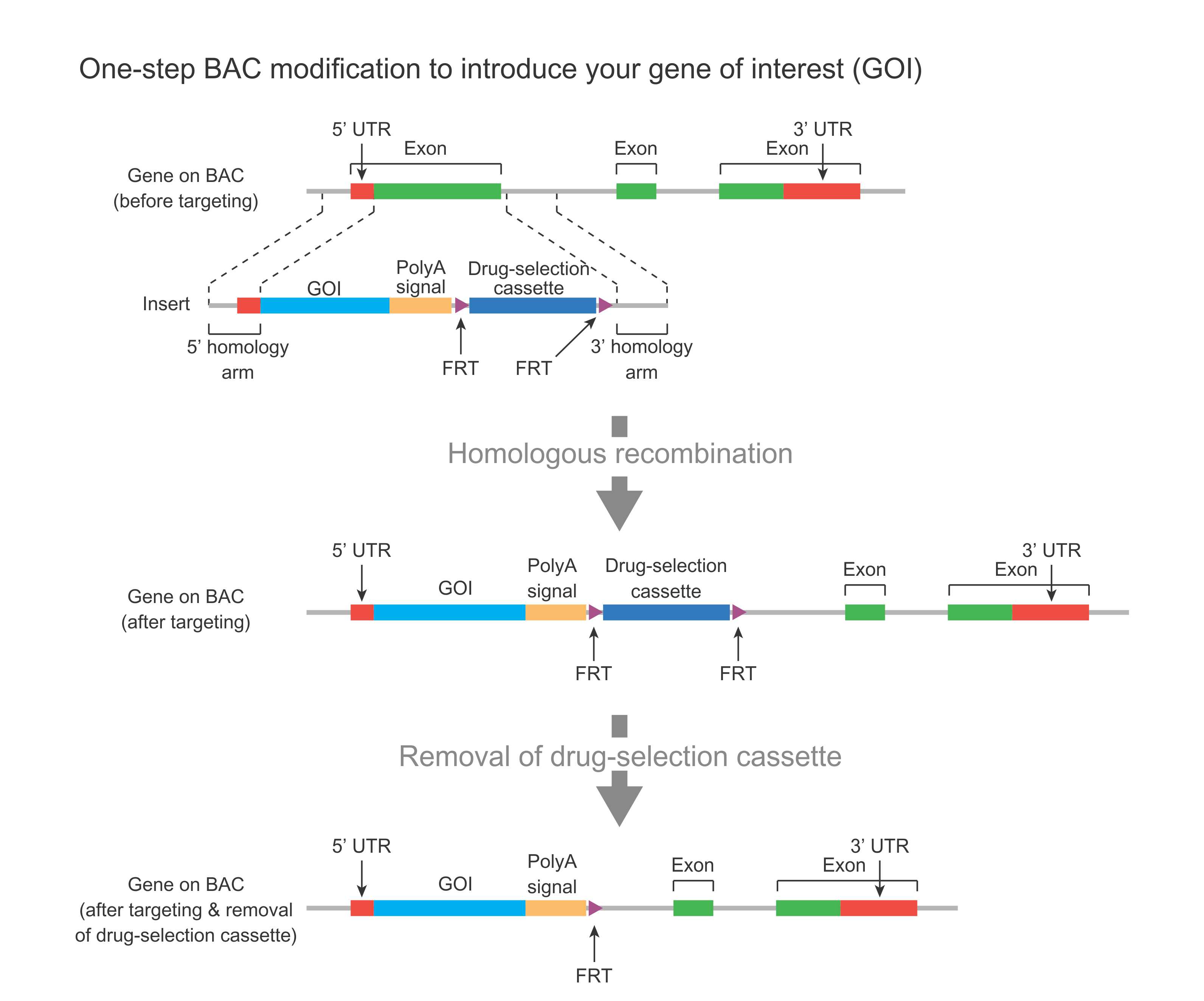
Figure 1. Schematic representation of one-step BAC modification for introducing a gene of interest (ROI) such lacZ or GFP reporter behind the promoter of a gene.
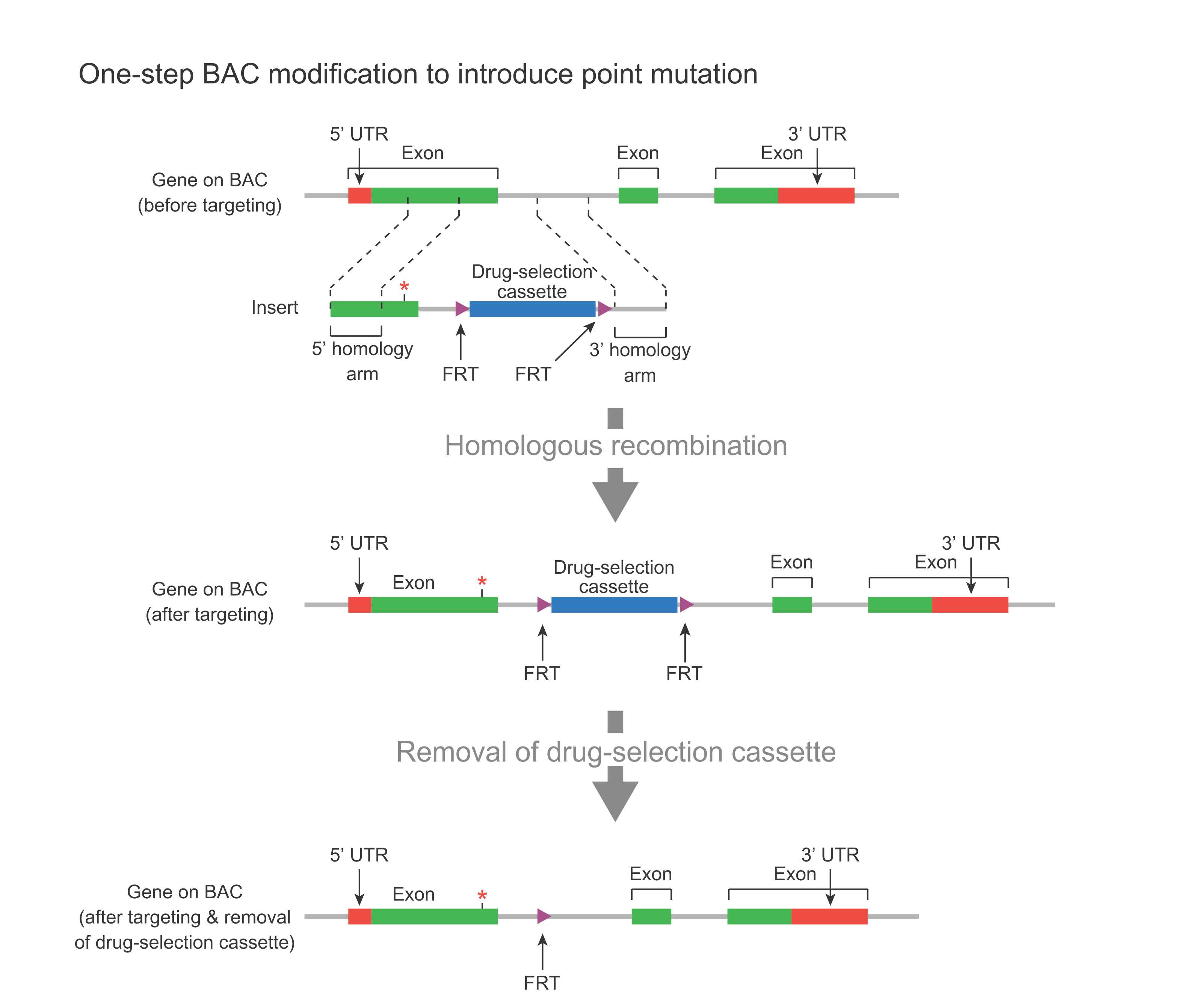
Figure 2. Schematic representation of one-step BAC modification for introducing a point mutation in the first exon of a gene along with an FRT scar sequence in the downstream intron.
Two-step BAC modification
The two-step modification method is used to introduce minor changes such as point mutations into a gene on the BAC without leaving behind the FRT scar sequence (Figure 3). This approach is preferred over the one-step approach when there is no place in the vicinity of the desired mutation to leave behind the Frt scar sequence. For example, if the gene to be modified is a very large single-exon gene, and the desired point mutation is in the middle of the gene, then there is no good place near the point mutation to place the scar sequence. In this case, the two-step method can be used. This method relies on positive and negative selection using a suitable drug-selection cassette (Figure 3). In the first step, this cassette is used to replace the target region via homologous recombination, which is facilitated by positive selection. In the second step, the cassette is further replaced with the final sequence containing desired mutation via homologous recombination, which is facilitated by negative selection.
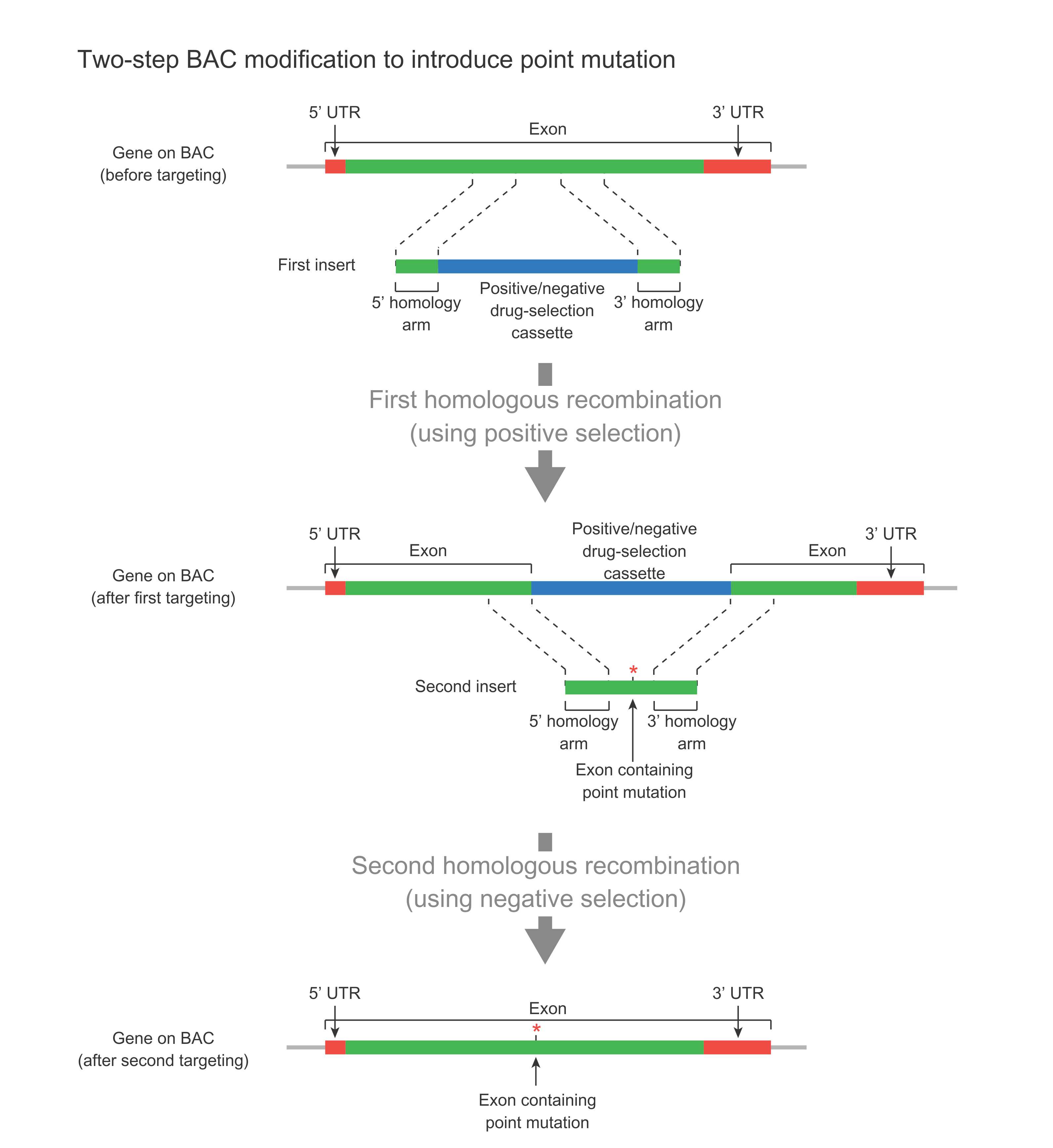
Figure 3. Schematic representation of two-step BAC modification for introducing a point mutation in a large single-exon gene without leaving behind any scar sequence.
Add drug-selection or visualization marker to BAC backbone
A drug-selection marker such as neomycin resistance or a visualization marker such as GFP can be added to the BAC backbone. When cells are transfected with such a modified BAC, the presence of a marker on the BAC backbone can facilitate the selection or visualization of cells successfully transfected with the BAC.
Transfer region of BAC onto plasmid
We can transfer any region of a BAC (up to 60 kb) onto a plasmid. This would allow the region to be studied without the influence of flanking sequences. For example, if an entire BAC is used to create transgenic mice in order to study the function of a gene on the BAC, the presence of other genes on the same BAC could confound the interpretation of the resulting phenotype. Isolating the gene onto a plasmid and making transgenic mice using the plasmid would circumvent this problem. An added benefit is the technical ease of working with plasmids versus BACs.
Validation of modified BAC
Modified region of the BAC will be confirmed by sequencing. Sometimes BACs can undergo deletions due to the presence of repetitive sequences. To mitigate this possibility, we will re-transform a BAC after it has been modified to obtain a single clone. We will then confirm that the clone has not undergone major deletions by performing PCR amplifications of multiple sites across the entire BAC.